Fan, K., Zhou, S., Jin, L. et al. BMC Immunol 24, 19 (2023). https://doi.org/10.1186/s12865-023-00556-1
Abstract
Background
Allergen-specific immunotherapy (AIT) is a causative treatment in allergic rhinitis (AR), comprising long-term allergen administration and over three years of treatment. This study is carried out for revealing the mechanisms and key genes of AIT in AR.
Methods
The present study utilized online Gene Expression Omnibus (GEO) microarray expression profiling dataset GSE37157 and GSE29521 to analyze the hub genes changes related to AIT in AR. Based on limma package, differential expression analysis for the two groups (samples of allergic patients prior to AIT and samples of allergic patients undergoing AIT) was performed to obtain differentially expressed genes (DEGs). Gene Ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis of DEGs were conducted using DAVID database. A Protein-Protein Interaction network (PPI) was built and a significant network module was acquired by using Cytoscape software (Cytoscape, 3.7.2).
Utilizing the miRWalk database, we identified potential gene biomarkers, constructed interaction networks of target genes and microRNAs (miRNAs) using Cytoscape software, and explore the cell type-specific expression patterns of these genes in peripheral blood using publicly available single-cell RNA sequencing data (GSE200107). Finally, we are using PCR to detect changes in the hub genes that are screened using the above method in peripheral blood before and after AIT treatment.Results
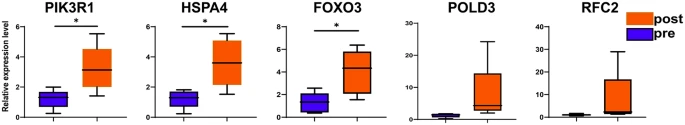 |
| Hub gene mRNA expression levels in patients before and after immunotherapy. *P < 0.05 |
Conclusion
Our analysis has identified novel gene signatures, thereby contributing to a more comprehensive understanding of the molecular mechanisms underlying AIT in the treatment of AR.
No comments:
Post a Comment