Background
Patients with chronic rhinosinusitis with nasal polyps (CRSwNP) and comorbid asthma have more severe disease and are difficult to treat. However, the molecular endotypes associated with CRSwNP with comorbid asthma (CRSwNP + AS) are not clear. This study aimed to investigate the characteristics of type 2 inflammation and the molecular signatures associated with CRSwNP + AS.
Methods
A total of 195 subjects; including 65 CRSwNP + AS patients, 99 CRSwNP-alone patients, and 31 healthy control subjects; were enrolled in the study. Nasal tissues from patients with CRSwNP + AS, CRSwNP-alone and control subjects were assessed for infiltration of inflammatory cells and concentrations of total IgE. Whole-transcriptome sequencing was performed and differentially expressed (DE) mRNAs and long non-coding RNAs (lncRNAs) and their associated pathways were analyzed. The correlations between type 2 cytokines and local eosinophils, tissue IgE, and transcriptome signatures were evaluated.
Results
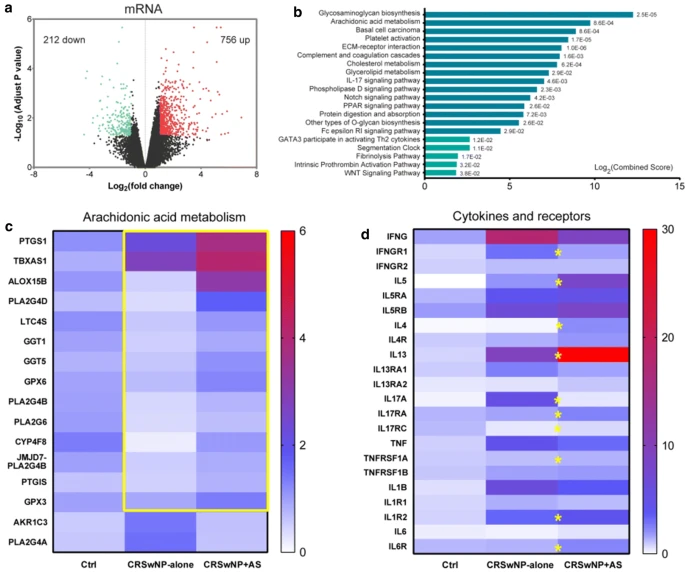 |
| Differentially expressed genes and pathways between CRSwNP + AS
and CRSwNP-alone. a Volcano plots illustrating DE-mRNAs of
CRSwNP + AS versus CRSwNP-alone identified by RNA sequencing.
b Top 15 KEGG pathways (blue column) and top 5 BioCarta pathways
(turquoise column) significantly enriched by DE-mRNAs.
c The expression of arachidonic acid metabolism-related DE-mRNAs
between CRSwNP + AS and CRSwNP-alone.
The colour coding of heat maps represents the gene expression
level normalized to Control group, calculated based on fragments
per kilo-base of exon per million fragments mapped (FPKM).
Yellow box indicates the up-regulated genes in CRSwNP + AS group.
d The expression of critical cytokines and their receptors that indicated
the activity of different inflammatory endotypes.
Yellow stars represent significantly differentially expressed
genes between CRSwNP + AS and CRSwNP-alone.
P < 0.05 were considered statistically significant. CRSwNP chronic rhinosinusitis
with nasal polyps, AS asthma, DE differentially expressed,
KEGG Kyoto Encyclopedia of Genes and Genomes |
|
Significantly higher local eosinophil infiltration and higher levels of total IgE were found in nasal tissues from CRSwNP + AS patients than in nasal tissues from CRSwNP-alone patients. Furthermore, atopy and recurrence were significantly more frequent in patients with CRSwNP + AS than in patients with CRSwNP-alone (62.5% vs 28.6% and 66.7% vs 26.9%, respectively). RNA sequencing analysis identified 1988 common DE-mRNAs, and 176 common DE-lncRNAs shared by CRSwNP + AS versus control and CRSwNP-alone versus control. Weighted gene coexpression network analysis (WGCNA) identified LINC01146 as hub lncRNA dysregulated in both subtypes of CRSwNP. Overall, 968 DE-mRNAs and 312 DE-lncRNAs were identified between CRSwNP + AS and CRSwNP-alone. Both pathway enrichment analysis and WGCNA indicated that the phenotypic traits of CRSwNP + AS were mainly associated with higher activities of arachidonic acid metabolism, type 2 cytokines related pathway and fibrinolysis pathway, and lower activity of IL-17 signalling pathway. Furthermore, the expression of type 2 cytokines; IL5 and IL13, was positively correlated with local eosinophil infiltration, tissue IgE level, and the expression of DE-mRNAs that related to arachidonic acid metabolism. Moreover, WGCNA identified HK3-006 as hub lncRNA in yellow module that most positively correlated with phenotypic traits of CRSwNP + AS.
Conclusions
Patients with CRSwNP + AS have distinct type 2-high inflammation-associated molecular signatures in nasal tissues compared to patients with CRSwNP-alone.